-Towards improving the citrus breeding efficiency -
The whole genome sequence of Satsuma mandarin, one of the major early maturing citrus variety in Japan was decoded. About 29,000 genes including 91 genes involved in coloring and fruit-setting of citrus fruits were deduced. The genome sequence information will facilitate for improving the efficiency in the citrus breeding program, productivity, and quality of citrus fruits.
Overview
- The National Agriculture and Food Research Organization, in collaboration with the National Institute of Genetics, decoded the entire genome sequence of Satsuma mandarin (Miyagawa Wase). The obtained genome size was approximately 360 million base pairs.
- A gene modeling analysis deduced about 29,000 genes in the genome. A gene annotation analysis identified 91 genes for the biosynthesis of carotenoids that is involved in fruit coloring, as well as genes for gibberellin biosynthesis that regulates fruit setting.
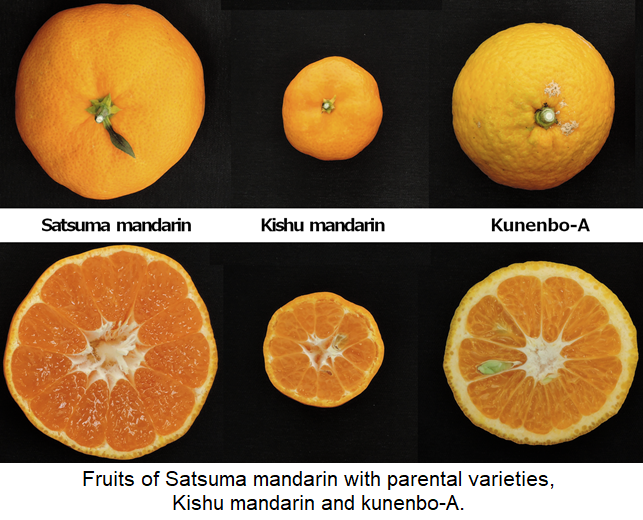
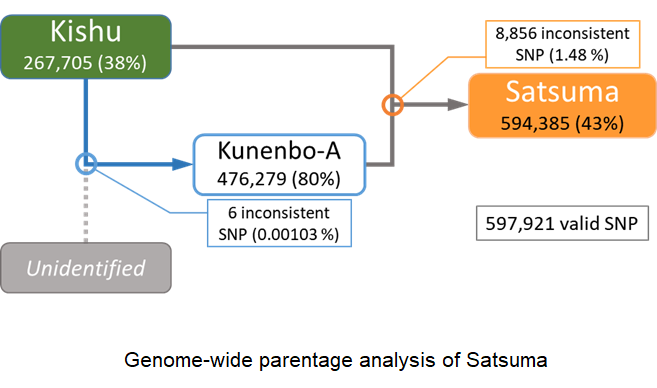
- Whole genome sequencing analysis of Kishu mandarin and kunenbo-A, which are the parents of Satsuma mandarin, verified the parentage of kunenbo-A as an offspring of Kishu mandarin. Consequently, Satsuma mandarin is confirmed to be a back-crossed progeny of Kishu mandarin.
- The obtained genome sequence information will be applied to the genome-wide association study for accelerating the identification and functional analysis of pivotal genes that regulate productivity and fruit traits of citrus. It will contribute to improving the breeding efficiency of citrus.
- These results were published in the international scientific journal Frontiers in Genetics on December 5, 2017.
For Inquiries
Contact: http://www.naro.affrc.go.jp/english/inquiry/index.html
Publication
Tokurou Shimizu, Yasuhiro Tanizawa, Takako Mochizuki, Hideki Nagasaki, Terutaka Yoshioka, Atsushi Toyoda, Asao Fujiyama, Eli Kaminuma, Yasukazu Nakamura (2017) Draft sequencing of the heterozygous diploid genome of Satsuma (Citrus unshiu Marc.) using a hybrid assembly approach. Frontiers in Genetics. 8:180, doi: 10.3389/fgene.2017.00180
Supplement