NARO
Saga University
The National Agricultural Research Organization (NARO) in collaboration with Saga University has discovered a new virus defense mechanism in plants from soybean. In order to escape the immune system of a host organism, certain viruses create a "hiding location" in infected cells where they amplify the genome. Rsv4 protein made from soybean mosaic virus resistance gene (Rsv4) can prevent infection by finding the "Hiding location" of the virus and digesting the viral genome. Furthermore, by applying this mechanism, we succeeded in creating artificial proteins that suppress the growth of various viruses. This research result helps to develop soybean varieties resistant to soybean mosaic virus, also it can be expected to lead to new control methods against viral diseases in various crops.
Overview
Soybean mosaic virus (SMV) infects soybean and reduces the yield and quality. Therefore, SMV-resistant soybean varieties have been cultivated as a means of control since ancient times. However, in recent years, SMV mutant strains which is not effective to conventional SMV resistant gene has emerged and has become an issue.
Research team from NARO and Saga University has identified soybean SMV resistance Rsv4 gene effective against a wide range of SMV including mutants and clarified that the Rsv4 protein produced from the gene prevents SMV infection by a completely new mechanism that has not been known so far. RNA viruses such as SMV are known to create a "hiding location" in the cell and replicate the genome there, so that they cannot be found in the defense mechanism of the infected organism. The Rsv4 protein was found to prevent viral infection by finding the "hiding location" and digesting the viral genome.
The research team applied this mechanism to various virus that affects crops other than soybeans such as Tomato mosaic virus of tomato, Cucumber mosaic virus which affects many plants etc., and succeeded at the laboratory level to create artificial proteins that digest virus genome by sneaking into the "hiding location" of these viruses and suppressing their growth.
This result will enable the development of resistance varieties that are difficult to break down by integrating the Rsv4 gene into soybean varieties with conventional resistance genes. In addition, it is expected that new resistance genes can be designed against viruses that cannot use natural resistance genes or mutant viruses that break resistance genes. The results will be published in the online version of the international science journal Nature Communications (September 27, 2019 issue).
Publication
Ishibashi K, Saruta M, Shimizu T, Shu M, Anai T, Komatsu K, Yamada N, Katayose Y, Ishikawa M, Ishimoto M, Kaga A. Soybean antiviral immunity conferred by dsRNase targets the viral replication complex. Nature Communications DOI:10.1038/s41467-019-12052-5.
For Inquiries
Contact: http://www.naro.go.jp/english/inquiry/index.html
Reference Information

Photo 1 Upper leaves (right) of soybean which is affected by soybean mosaic virus widely and shows mosaic symptoms. The color of the leaf part where viruses are accumulated much will become lighter.
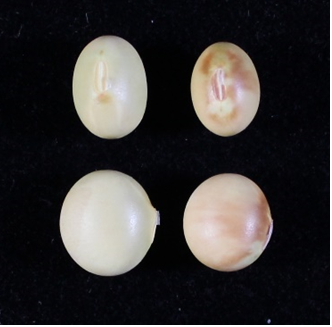
Photo 2 Seeds harvested from soybean inoculated with soybean mosaic virus
Brown mottled seeds (right) produced in the soybean cultivar "Enrei", seeds of the line in which Rsv4 gene was introduced into "Enrei" by crossing and DNA marker selection (left).
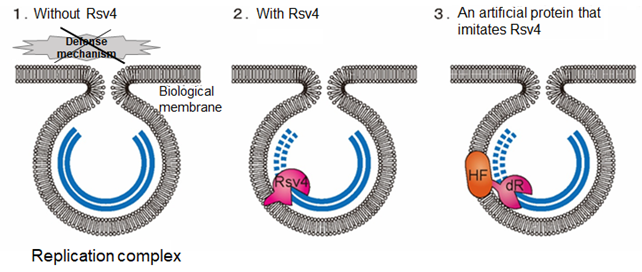
Fig. 1 (1) When there is no Rsv4. The viral genome (blue) hides from the defense mechanism in infected cells and replicates.
(2) When there is Rsv4. Rsv4 enters the replication complex and degrades the viral genome (double-stranded RNA).
(3) A new virus control method using an artificial protein that mimics/imitates Rsv4. By fusing the protein (HF) used for virus replication and double-stranded RNase (dR), double-stranded RNase can be sent into the replication complex.




